Research
Selected grants
Main research projects
GPCR Modulation
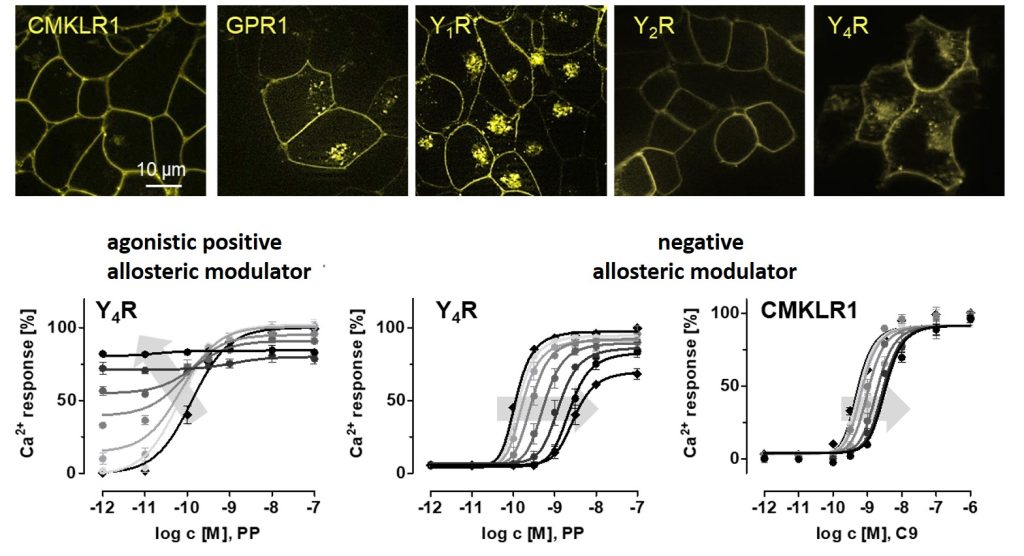
G protein-coupled receptors are targets of a variety of biologically relevant substances, such as neuropeptides and hormones. The receptor response after ligand stimulation can be modulated by small molecule substances acting at an allosteric binding site. This modulation may lead to increasing or decreasing affinity or efficacy of the native ligand, to enhancing or attenuating the G protein-dependent receptor response, or by effecting the recruitment of intracellular adapter proteins. Thus, a variety of intracellular signaling pathways can be targeted with modulators that are temporally and spatially dependent on the natural ligand. The aim of this project is to understand the underlying mechanisms of receptor activation and the modulation of these mechanisms by allosteric modulators, and thus to identify new concepts for the development of therapeutically relevant molecules.
GPCR Structure and Dynamics
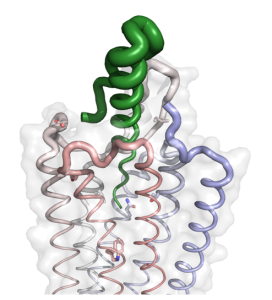
G protein-coupled receptors (GPCRs) can recognize a wide variety of ligands, and interact with many cellular components. This is enabled by a remarkable conformational flexibility. We are interested in how GPCRs recognize their interaction partners with an emphasis on how selectivity and activity of (endogenous) peptide ligands is achieved. We hypothesize that specificity is coordinated by key amino acids in the binding pockets as well as by the dynamic interactions between peptide and receptor. We investigate these questions by mutagenesis and functional assays, and collaborate intensively with structural biology labs. The picture on the right shows the cryo EM structure of the human neuropeptide Y2 receptor in complex with its endogenous peptide ligand neuropeptide Y (green). The thickness of the ribbon represents the residual flexibility of the protein even with the ligand bound. Important residue of the binding pocket are highlighted as sticks.
Shuttling - Peptide Mediated Cell Specific Drug Delivery
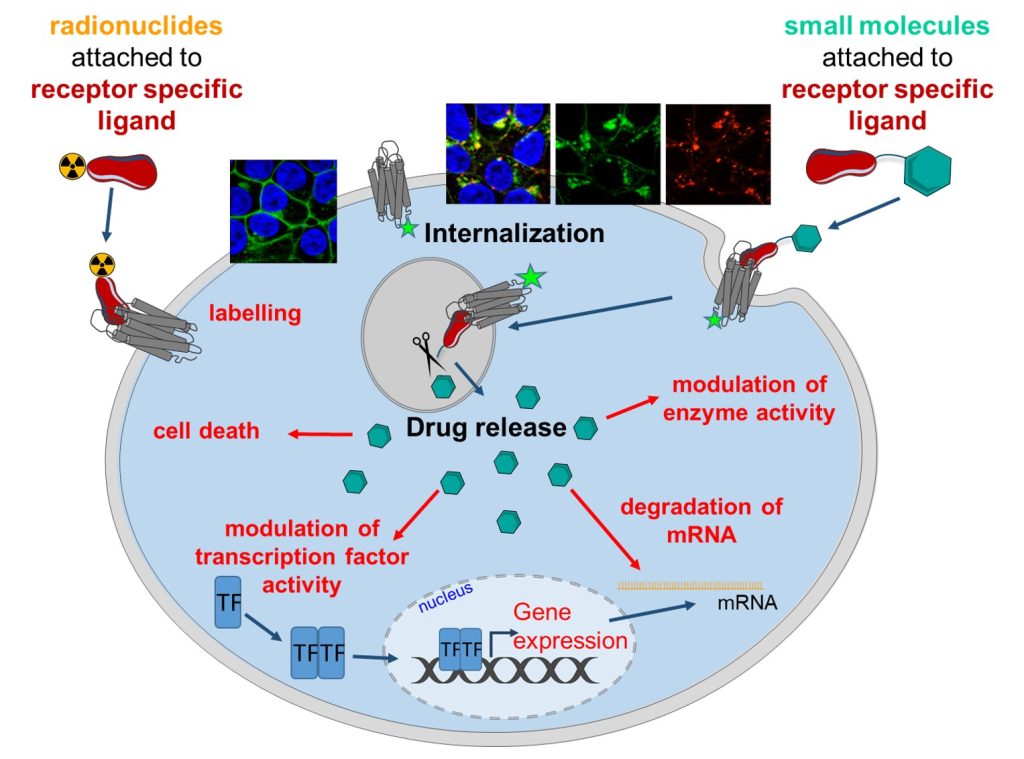
Side effects, also known as adverse reactions, represent a major hurdle in drug development. New technologies improving drug delivery and accumulation in specific target cells while reducing off-target effects and decreasing unwanted toxicities play a major role in the translation of potential excellent drug candidates to clinical studies and approval. In our projects we exploit the cell specific surface expression of G protein-coupled receptors (GPCR), e.g. highly restricted endogenous receptor expression or overexpression in cancer cells, to develop shuttling systems mediating selective drug uptake. We modify endogenous peptide GPCR-ligands to increase their specificity and stability and subsequently link them to small chemical drug molecules or RNA therapeutics by cleavable or stable linkers. Binding of the peptide-drug-conjugates to the GPCR will result in specific receptor mediated uptake and delivery of the drug to its intracellular target sites. Besides the modulation of cellular phenotypes, radioactive or cytotoxic molecules are of great interest for cancer imaging or therapy.
Therapeutic Peptides and Proteins
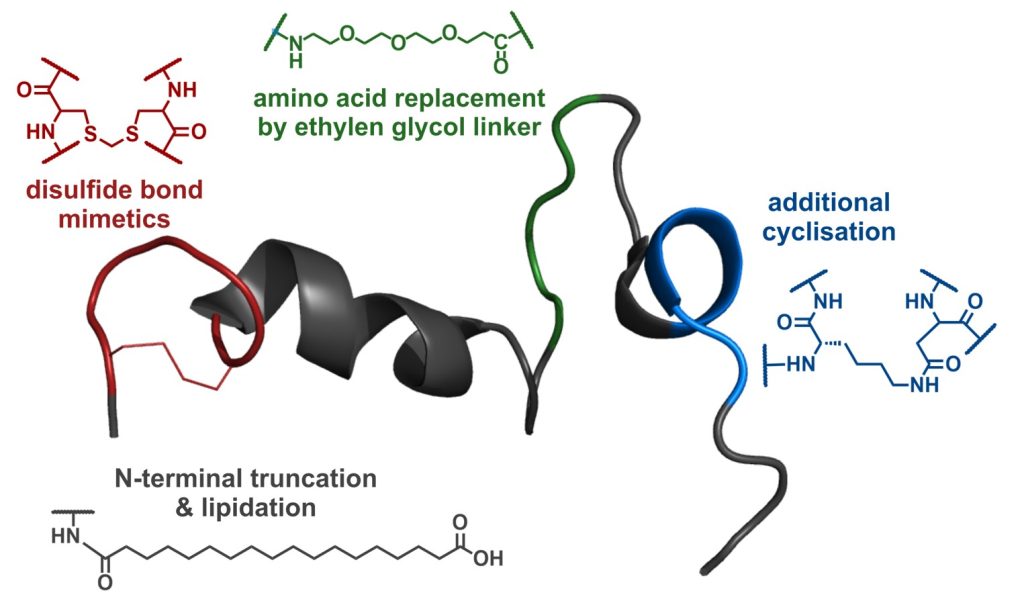
Therapeutic peptides and proteins have gained increasing attention in recent years as they offer advantages over traditional small molecule drugs. Especially their high target specificity, selectivity, and predictable metabolites are attractive features. In our lab, we combine chemical methods for the synthesis of modified or constrained peptides with their biological characterization to transform naturally occurring molecules into therapeutic agents.
Taking into account knowledge of binding modes gained by computational or experimental methods, we have successfully optimized several therapeutically relevant peptides regarding stability in blood plasma and biological activity at their target receptors. Our projects focus on the development of therapeutic peptides and proteins targeting cancer, obesity and related diseases, and the regulation of blood pressure.
Biomaterials – From Modified Peptides and Proteins to Selective Immobilization of Molecules and Cells
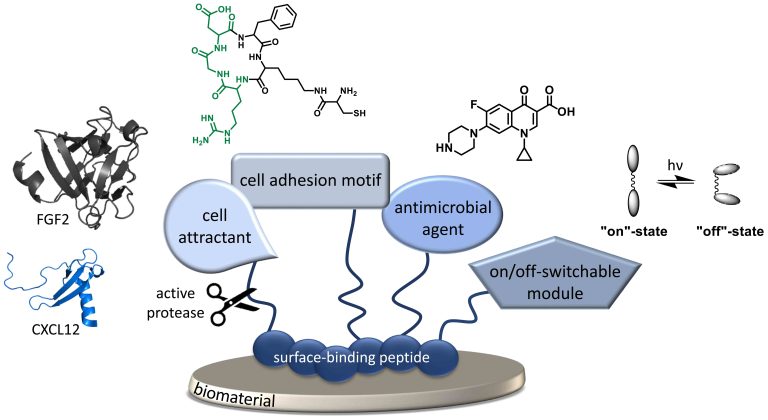
Novel biomaterials have been discovered that provide some “smart” functions, which includes a combination of biodegradability with strength and elasticity without undesired host responses. However, all biomaterials are prone to foreign body reactions resulting in inflammation and encapsulation of the device. To increase on the one hand cell viability as well as tissue integration and on the other hand to reduce infection risk and implant failure, it is desirable to coat such materials with bio-functional peptides addressing the device-contacting tissue. Moreover, smart biomaterials might induce the guidance to injured tissue sites of desired cell types, which can be realized by stable gradients of e.g. chemokines. In this project, a surface-binding peptide acts an anchor unit on several materials and is decorated with diverse modular functionalities like ECM-derived cell adhesive RGD-motifs or GAG-binding peptides, antibiotics or antimicrobial peptides as well as protease-releasable chemokines/cytokines and growth factors like CXCL12 or FGF2, e.g. to decoy desired cell types towards the biomaterial interface. These peptides and proteins are synthesized and the immobilization on different materials as well as the cellular response are characterized.
Besides the medical application of functional materials, these are already important for catalytic purposes. Enzymes are gaining importance in the chemical industry and for biotechnological applications, e.g. for the synthesis of fatty acids or steroid derivatives. They can perform complex chemical reactions without high temperature, high pressure, or rare and expensive catalysts. Immobilization can lead to stabilization of the protein, higher activity due to oriented surface presentation as well as easy separation of enzyme and product. In this project, specific linkers enable the electron-driven activation of the enzyme cytochrome P450 without the need of co-factors, solely by applying a defined current flow. Moreover, by incorporating azo photo-switches the linker can be used to distinguish between an “on”- and “off”-state of the enzyme.