Z4 – Platform technologies for the analysis of glycosaminoglycans and mediator proteins
Within the scope of subproject Z4, qualitative and quantitative protein analysis is carried out as a service for all Transregio 67 projects. For this purpose, the development of methods for the highly sensitive detection of low molecular weight proteins as well as highly modified proteins, including cytokines and growth factors, is being developed.
- The identification and quantification of proteins extracted by microdialysis from injured tissue during the wound healing process by means of nano-HPLC / nano-ESI mass spectrometry
- Protein expression analysis from cell models
- Enrichment and determination of glycosylation
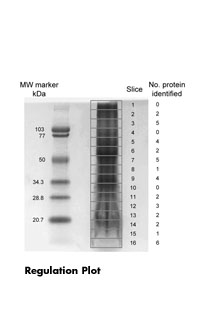
- Protein separation by 1D / 2D-SDS-PAGE or HPLC (size exclusion chromatography, C4 reverse phase chromatography)
- Implementation of trainings / introduction of TRR67 employees.
Publications
- Großkopf H, Vogel S, Müller CD, Köhling S, Dürig JN, Möller S, Schnabelrauch M, Rademann J, Hempel U, von Bergen M, Schubert K. Identification of intracellular glycosaminoglycan-interacting proteins by affinity purification mass spectrometry. Biological Chemistry, vol. 402, no. 11, 2021, pp. 1427-1440. https://doi.org/10.1515/hsz-2021-0167.
- Kratochvil I, Hofmann T, Rother S, Schlichting R, Moretti R, Scharnweber D, Hintze V, Escher BI, Meiler J, Kalkhof S, von Bergen M. Mono(2-ethylhexyl) phthalate (MEHP) and mono(2-ethyl-5-oxohexyl) phthalate (MEOHP) but not di(2-ethylhexyl) phthalate (DEHP) bind productively to the peroxisome proliferator-activated receptor γ. Rapid Commun Mass Spectrom. 2019 May;33 Suppl 1:75-85. doi: 10.1002/rcm.8258. Epub 2019 Jan 1.
- Schmidt JR, Vogel S, Moeller S, Kalkhof S, Schubert K, von Bergen M, Hempel U. Sulfated hyaluronic acid and dexamethasone possess a synergistic potential in the differentiation of osteoblasts from human bone marrow stromal cells. J Cell Biochem 2019;120:8706-8722. doi:10.1002/jcb.28158
- Jouy F, Lohmann N, Wandel E, Ruiz-Gómez G, Pisabarro MT, Beck-Sickinger AG, Schnabelrauch M, Moeller S, Simon JC, Kalkhof S, von Bergen M, Franz S. Sulfated hyaluronan attenuates inflammatory signaling pathways in macrophages involving induction of antioxidants. Proteomics. 2017. [Epub ahead of print]
- Förster Y, Schmidt JR, Wissenbach DK, Pfeiffer SE, Baumann S, Hofbauer LC, von Bergen M, Kalkhof S, Rammelt S. Microdialysis Sampling from Wound Fluids Enables Quantitative Assessment of Cytokines, Proteins, and Metabolites Reveals Bone Defect-Specific Molecular Profiles. PLoS One. 2016;11:e0159580.
- Rother S, Samsonov SA, Hofmann T, Blaszkiewicz J, Kohling S, Moeller S, Schnabelrauch M, Rademann J, Kalkhof S, von Bergen M, Pisabarro MT, Scharnweber D, Hintze V. Structural and functional insights into the interaction of sulfated glycosaminoglycans with tissue inhibitor of metalloproteinase-3 – a possible regulatory role on extracellular matrix homeostasis. Acta Biomater. 2016; 45:143-54.
- Kalkhof S, Schildbach S, Blumert C, Horn F, von Bergen M, Labudde D. PIPINO: A Software Package to Facilitate the Identification of Protein-Protein Interactions from Affinity Puri-fication Mass Spectrometry Data. Biomed Res Int. 2016;2016:2891918
- Wewering F, Jouy F, Wissenbach DK, Gebauer S, Blüher M, Gebhardt R, Pirow R, von Bergen M, Kalkhof S, Luch A, Zellmer S. Characterization of chemical-induced sterile inflamma-tion in vitro: application of the model compound ketoconazole in a human hepatic co-culture system. Arch Toxicol. 2016. [Epub ahead of print]
- Schmidt JR, Kliemt S, Preissler C, Moeller S, von Bergen M, Hempel U, Kalkhof S. Osteoblast-released Matrix Vesicles, Regulation of Activity and Composition by Sulfa¬ted and Non-sulfated Glycosaminoglycans. Mol Cell Proteomics. 2016;15:558-72.
- Jouy F, Müller SA, Wagner J, Otto W, von Bergen M, Tomm JM. Integration of conventional quantitative and phospho-proteomics reveals new elements in activated Jurkat T-cell receptor pathway maintenance. Proteomics. 2015;15:25-33.
- Hofmann T, Fischer AW, Meiler J, Kalkhof S. Protein structure prediction guided by crosslinking restraints – A systematic evaluation of the impact of the crosslinking spacer length. Methods. 2015; pii: S1046-2023(15)00207-8.
- Hofmann T, Samsonov SA, Pichert A, Lemmnitzer K, Schiller J, Huster D, Pisabarro MT, von Bergen M, Kalkhof S. Structural analysis of the interleukin-8/glycosaminoglycan interactions by amide hydrogen/deuterium exchange mass spectrometry. Methods. 2015 2015;1:45-53.
- Jouy F, Müller SA, Wagner J, Otto W, von Bergen M, Tomm JM. Integration of conventional quantitative and phospho-proteomics reveals new elements in activated Jurkat T-cell receptor pathway maintenance. Proteomics. 2015; 15(1):25-33
- Kalkhof S, Förster Y, Schmidt J, Schulz MC, Baumann S, Weißflog A, Gao W, Hempel U, Eckelt U, Rammelt S, von Bergen M. Proteomics and metabolomics for in situ monitoring of wound healing. Biomed Res Int. 2014:934848.
- Blumert C, Kalkhof S, Brocke-Heidrich K, Kohajda T, von Bergen M, Horn F. Analysis of the STAT3 interactome using in-situ biotinylation and SILAC. J Proteomics. 2013 6:370-86.
- Kliemt S, Lange C, Otto W, Hintze V, Möller S, von Bergen M, Hempel U, Kalkhof S. Sulfated hyaluronan containing collagen matrices enhance cell-matrix-interaction, endocytosis, and osteogenic differentiation of human mesenchymal stromal cells. J Proteome Res. 2013;12:378-89.
- Möbius K, Nordsieck K, Pichert A, Samsonsov SA, Thomas L, Schiller J, Kalkhof S, Pisabarro MT, Beck-Sickinger AG, Huster D. Investigation of lysine side chain interactions of interleukin-8 with heparin and other glycosaminoglycans studied by a methylation-NMR approach. Glycobiology. 2013;23:1260-9.
- Müller SA, van der Smissen A, von Feilitzsch M, Anderegg U, Kalkhof S, von Bergen M. Quantitative proteomics reveals altered expression of extracellular matrix related of proteins of human primary dermal fibroblasts in response to sulphated hyaluronan and collagen applied as artificial extracellular matrix. J Mater Sci Mater Med. 2012;23:3053-65.
- Salbach J, Kliemt S, Rauner M, Rachner TD, Goettsch C, Kalkhof S, von Bergen M, Möller S, Schnabelrauch M, Hintze V, Scharnweber D, Hofbauer LC. The effect of the degree of sulfation of glycosaminoglycans on osteoclast function and signaling pathways. Biomaterials. 2012;33:8418-29.
- Goettsch C, Kliemt S, Sinnigen K, von Bergen M, Hofbauer LC, Kalkhof S. Quantitative proteomics reveals novel functions of osteoclast-associated receptor in STAT signaling and cell adhesion in human endothelial cells. J Mol Cell Cardiol. 2012;53:829-37.
- Abburi R, Kalkhof S, Oehme R, Kiontke A, Birkemeyer C. Artificats in amine analsis from anodic oxidation of organic solvents upon electrospray ionization for mass spectrometry. Eur J Mass Spectrom. 2012;18:301-12.
- Dautel F, Kalkhof S, Trump S, Michaelson J, Beyer A, Lehmann I, von Bergen M. DIGE based protein expression analysis of B[a]P-exposed hepatoma cells reveals a complex stress response including alterations in oxidative stress, cell cycle control, and cytoskeleton motility at toxic and subacute concentrations. J Proteome Res. 2011;10:379-93.
- Washietl S, Findeiss S, Müller SA, Kalkhof S, von Bergen M, Hofacker IL, Stadler PF, Goldman N. RNAcode: robust discrimination of coding and noncoding regions in comparative sequence data. RNA. 2011;17:578-94.
- Rockstroh M, Müller SA, Jende C, Kerzhner A, von Bergen M, Tomm JM. Cell frationation – an important tool for compartment proteomics. J Integr OMICS. 2011;1:135-43.
- Müller SA, Kohajda T, Findeiß S, Stadler PF, Washietl S, Kellis M, von Bergen M, Kalkhof S. Optimization of parameters for coverage of low molecular weight proteins. Anal Bioanal Chem. 2010;398:2867-81.
- Mörbt N, Mögel I, Kalkhof S, Feltens R, Röder-Stolinski C, Zheng J, Vogt C, Lehmann I, von Bergen M. Proteome changes in human bronchoalveolar cells following styrene exposure indicate involvement of oxidative stress in the molecular-response mechanism. Proteomics. 2009;21:4920-4933.
Contact

Prof. Dr. Martin von Bergen
Professor of Functional Proteomics
Helmholtz Institute for Environmental Research
Department of Molecular Systems Biology
Permoserstraße 15, 04318 Leipzig
Phone: +49 (0)341 235-1211
E-Mail: martin.vonbergen@ufz.de
Homepage: www.ufz.de/index.php?de=17634
